Translational Projects
Cell sorting is the process of separating a heterogeneous mixture of cells into several homogeneous fractions.
It can be used to identify rare, diseased cells present in a population of healthy cells.
Different cell types usually differ in their physical (e.g. size, stiffness, etc.) and chemical (e.g. response to
specific receptors) properties.
Cell sorters exploit this kind of inherent differences to separate two cell populations from each other.
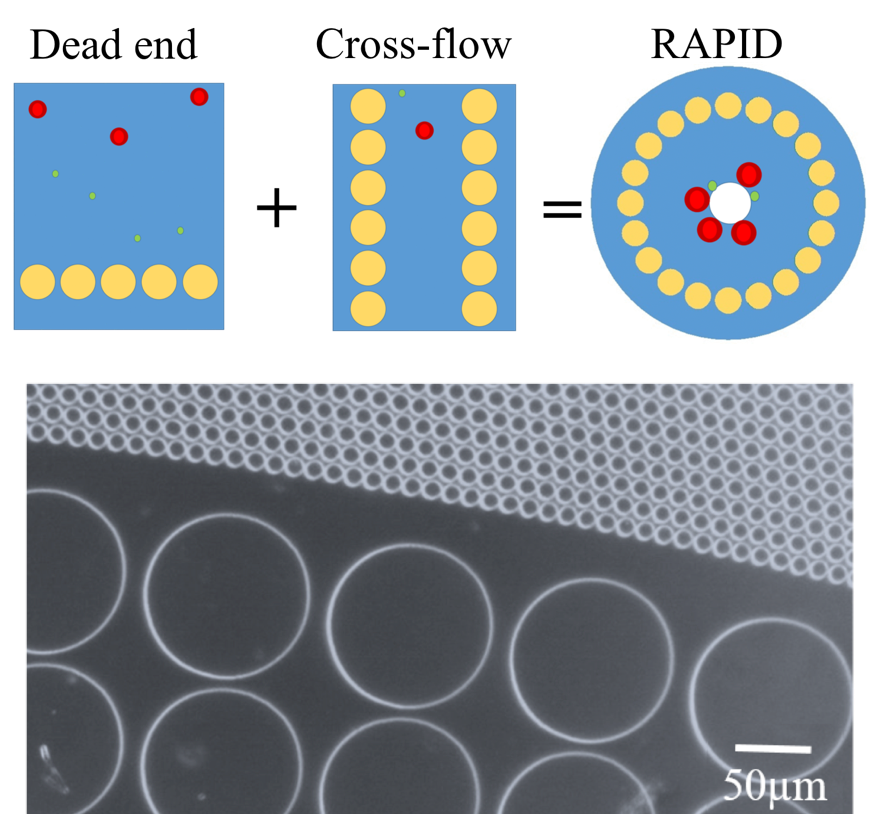
In our lab, we developed a microfluidic radial pillar device (RAPID) to fractionate blood cells based on the difference
in their biophysical properties, such as, size and deformability. This platform combines the selectivity of dead-end pillar design with the clog-free operation of cross-flow design, while keeping a small device footprint.
We operated RAPID with whole blood for close to 5 hours without clogging. Our current research is focussed on adapting this device device for enrichment of circulating tumour cells, and building a centrifugal portable prototype to replace pumps. For more details on RAPID, see our publications.
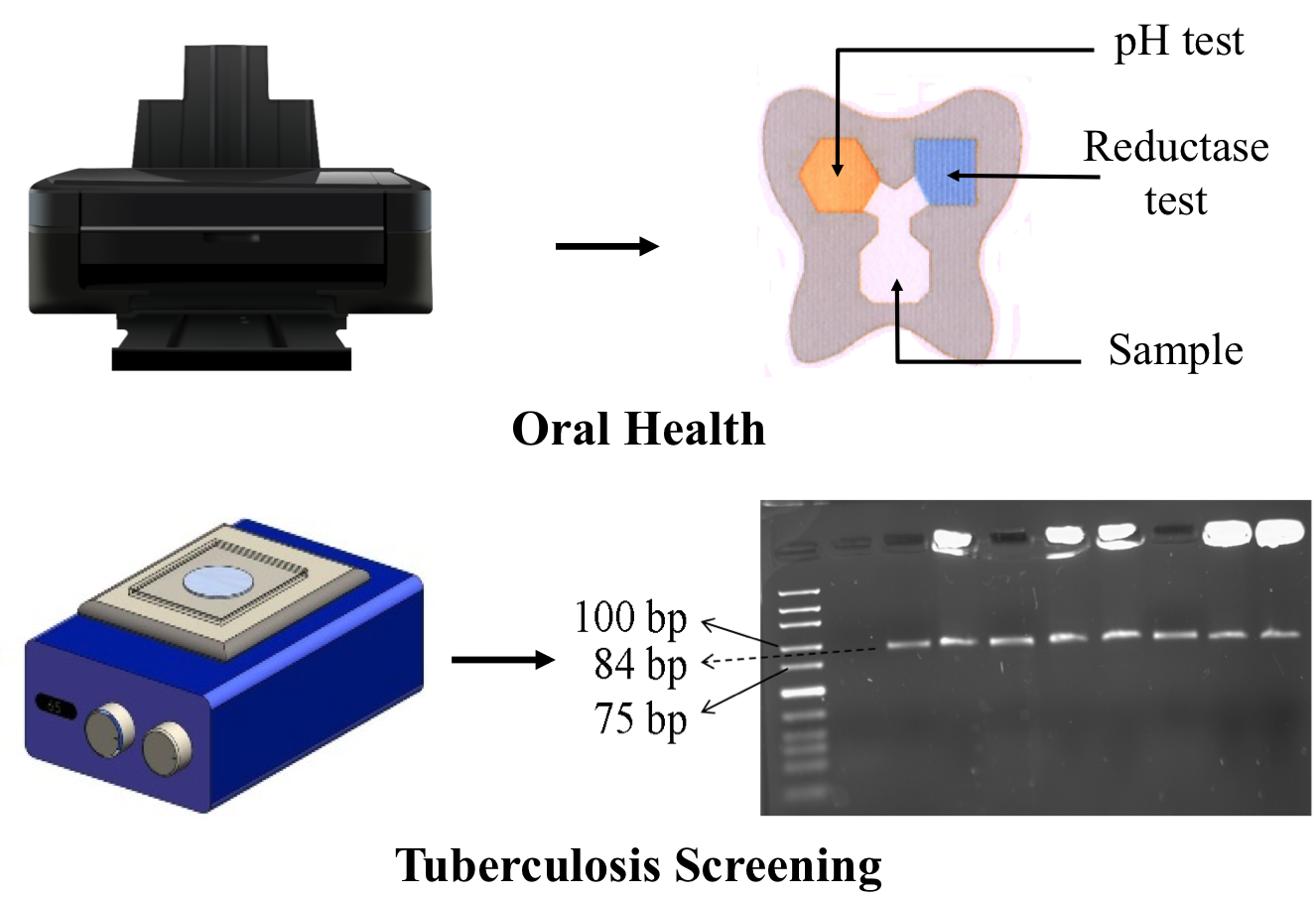
Paperfluidics is an upcoming field that uses paper as a substrate to develop microfluidic platforms, primarily for diagnostic applications for extreme resource-limited settings. We are interested in developing fully printable assays for various healthcare and environmental applications. We have recently reported a technique to pattern hydrophobic zones on paper by dispensing oil through a modified inkjet printer. One of the printed assays developed in our lab is a chip for home-based monitoring of oral health. In another project, we are developing a DNA-based screening assay for tuberculosis. Our platform works on the principle of isothermal amplification of DNA and obviates the need for an expensive thermocycler. We have managed to amplify purified tuberculosis DNA on a paper substrate in 10 minutes. For more details, see the publications and the related patents.
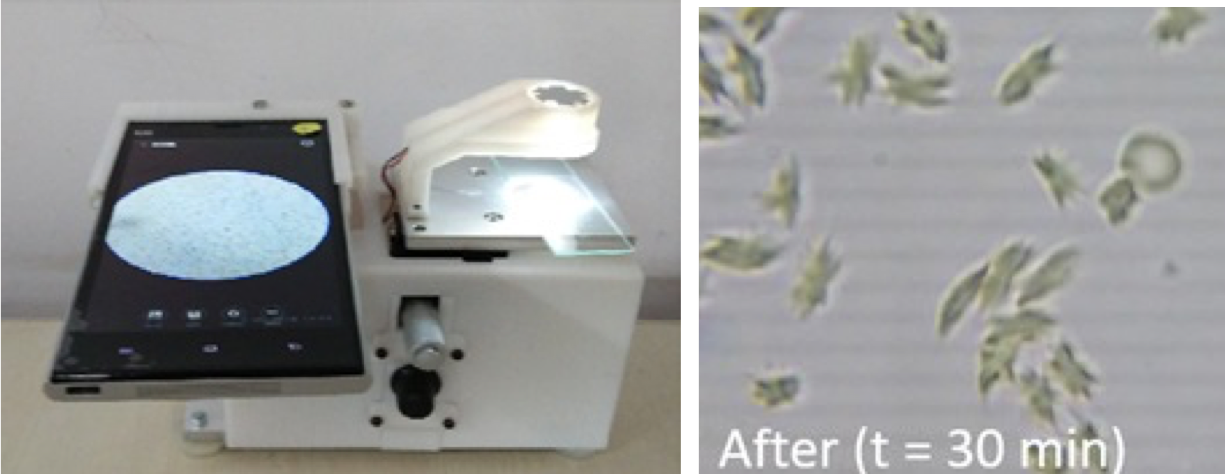

Sickle cell disease (SCD) is an inherited form of blood disorder due the presence of an abnormal form of hemoglobin (HbS). This results in polymerization of hemoglobin, particularly under hypoxic conditions, which makes the red blood cells (RBCs) less deformable and sickle-shaped. While there is no cure for the disease yet, early diagnosis can help in better management of the symptoms. Many areas in India, where this condition is prevalent, lack the facilities for a confirmed diagnosis of sickle cell disease.
We developed a platform combining blood chemistry, microfluidics, mobile phone microscopy and image processing
for automated diagnosis of sickle cell disease at the point-of-care. We optimised the oxygen scavenger concentration
and the temperature to vary the rate of polymerization of the mutated hemoglobin and the resulting cell shape. Unlike
most image processing methods, our technique can distinguish between sickle cell trait (heterozygous) and sickle cell disease (homozygous) samples in a single test. We reported some preliminary results in the proceedings
of the 20th International Conference on Miniaturized Systems for Chemistry and Life Sciences (MicroTAS 2016).
Along with the sickle cell assay, we also built a mobile phone microscope to take the diagnosis into the field. The microscope
design has been being taken up for commercialization by Medprime Technologies, an IIT Bombay start up company co-founded
by our lab alumnus Samrat.
Biological Physics
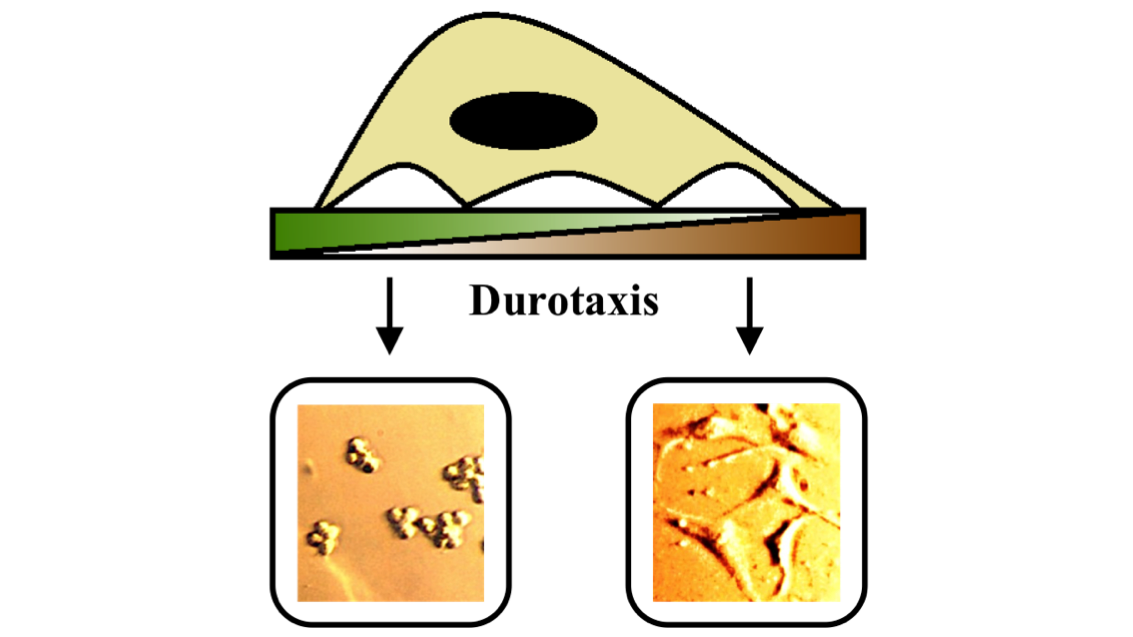

The behaviour of cells is strongly influenced by their environment. Unlike experiments on bulk cell populations, microfluidic devices allow us to control the local environment of single cells. We are interested in understanding how the physical properties of the cellular environment, such as, confinement, substrate stiffness, etc. affect the way cells move, change their shape and divide. This project is in collaboration with Dr. Shamik Sen, Department of Biosciences and Bioengineering, IIT Bombay.
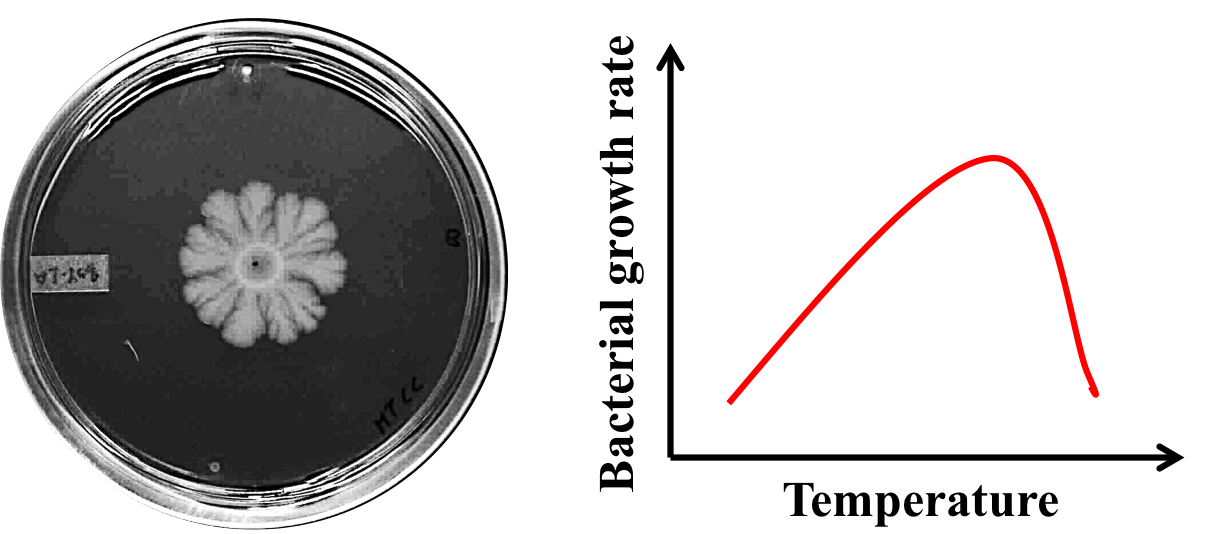

We are exploring how various physical parameters, such as, presence of nutrients and temperature affect the collective behaviour of motile and non-motile bacteria. The Paul lab built an experimental set-up for simultaneous automated imaging of four bacterial plates over several days. The entire imaging set-up can be housed inside a bacterial incubator. The images of bacterial colonies are then analysed to find the velocity of the bacterial front to understand the mechanism of colony expansion at different temperatures. The experiments done at IIT Bombay are then compared with simulations done by the group of Prasad Perlekar (Centre for Interdisciplinary Sciences, TIFR Hyderabad) to explain the behaviour of bacteria.
