Synthetic Biology
Designing Transcription factor (TF) decoys for regulating gene expression using DNA Nanotechnology
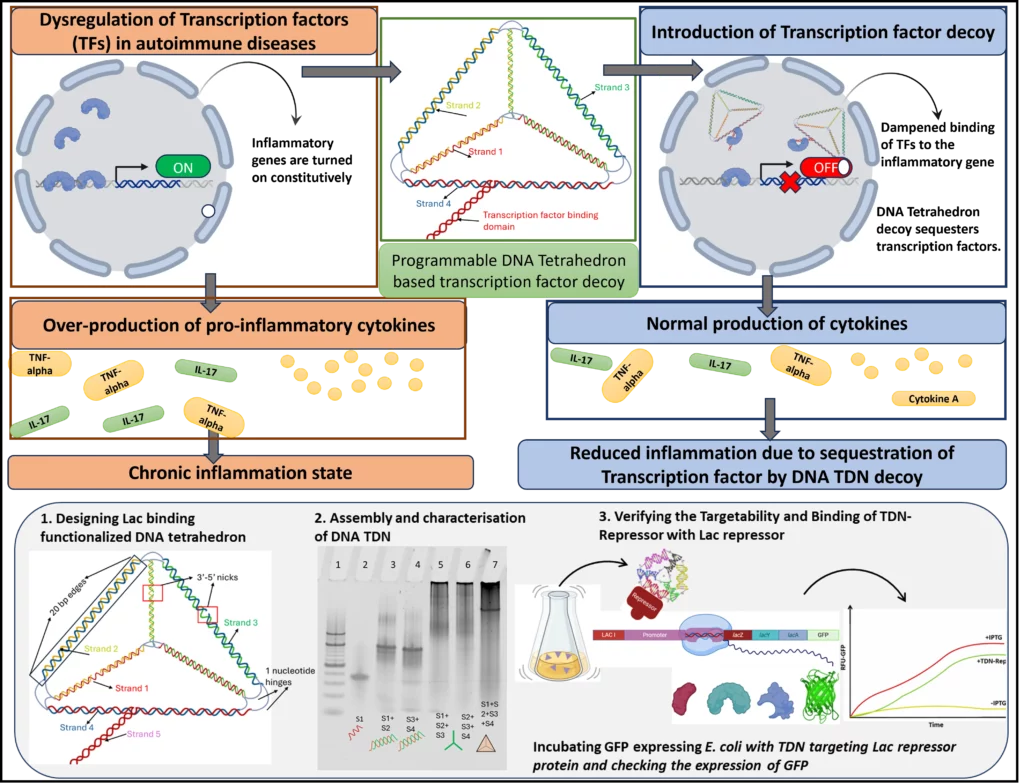
TF decoys are short ds DNA sequences that can act like a sponge to soak up free transcription factors in the cell and divert them from their native binding site. This can be harnessed to alter the protein expression and regulate immunological pathways. However, transfecting unmodified transcription factors (TFs) into cells is challenging because the negative charge of DNA causes repulsion at the cell membrane. Additionally, if they do enter, these oligonucleotides are rapidly degraded by nucleases and other enzymes within the cell. This issue can be solved by using DNA-based nanostructures. DNA can fold into different polyhedra due to its flexible nature and the turns of double-stranded DNA, which are also quite stable in the cellular environment, unlike double-stranded DNA-based circuits.
We are currently focused on building a DNA-based Tetrahedron that can mimic transcription factor binding sites, functioning as a decoy that can bind to transcription factors and regulate gene expression. We are currently testing the transfection and interaction of the TF decoy in E coli, which binds to the Lac repressor, and can regulate GFP expression.
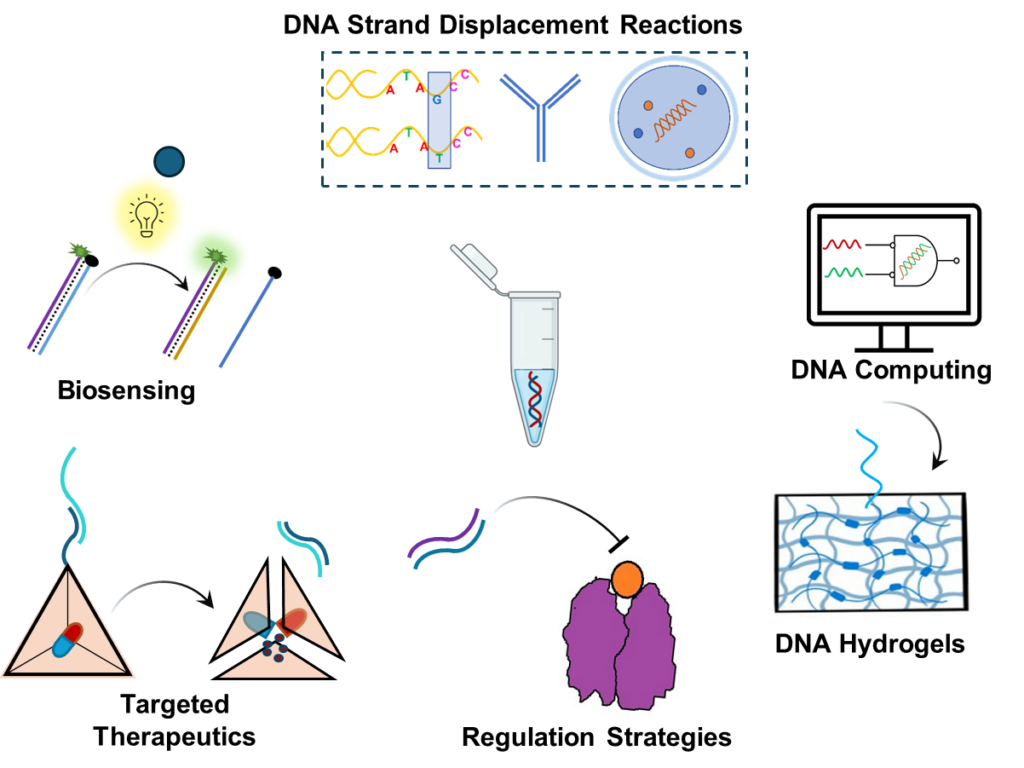
Applications of toehold mediated strand displacement reaction for biomedical applications
Dynamic DNA nanotechnology is applied in several biomedical applications, including biosensing for rapid pathogen detection and SNP sensing for personalized medicine. Additionally, these DNA constructs are utilized in therapeutics for controlled drug delivery, achieved by integrating DNA circuits with hydrogels to facilitate targeted drug release. This research aims to enhance the functionality of synthetic DNA in diagnostics and therapeutics, enabling innovative strategies to address significant health challenges.
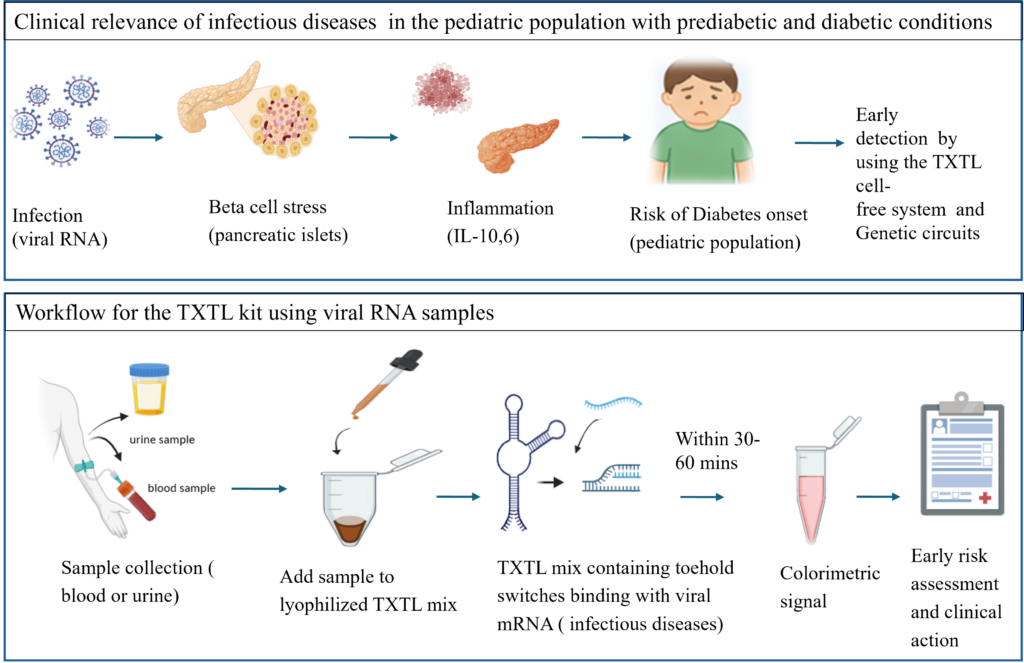
Cell-Free TXTL-Based Platform for Early Detection of Infection-Induced Diabetes Risk in Pediatric Populations
Infectious diseases can induce metabolic stress and contribute to pancreatic β-cell dysfunction in pediatric prediabetic and diabetic populations. This study presents a cell-free transcription–translation (TXTL)–based diagnostic platform for the rapid detection of infection-associated metabolic risk. The system integrates viral RNA sensing with biomarker-responsive genetic circuits to enable early identification of inflammatory responses and islet stress. A streamlined workflow utilizing blood or urine samples activates a lyophilized TXTL system, generating a colorimetric readout within 30–60 minutes. This portable and scalable platform facilitates early risk assessment and supports timely clinical intervention, particularly in resource-limited areas.
